Language / Ngôn ngữ:
McKaizer Institute — Longevity & Wellness Science
Discover how ultra-high-throughput single-cell technologies reveal why tissues age differently and what this means for extending healthy lifespan.
15 million cells analyzed
Dr. Cao’s lab mapped aging across 15 million individual cells revealing unprecedented tissue-specific aging patterns
Table of Contents
- The Revolution in Understanding Aging at Single Cell Resolution
- How Single Cell Sequencing Decodes the Biology of Aging
- Ultra High Throughput Methods That Map Millions of Aging Cells
- Discovering Senescent Cells Hidden Within Aging Tissues
- Nutritional and Lifestyle Factors That Influence Cellular Aging Trajectories
- From Cellular Maps to Clinical Interventions for Healthy Aging
- Novel Biomarkers Emerging From Single Cell Aging Atlases
- The Future of Personalized Longevity Through Cellular Analysis
- Frequently Asked Questions (20)
The Revolution in Understanding Aging at Single Cell Resolution

The Revolution in Understanding Aging at Single Cell Resolution
For decades, scientists studied aging like astronomers studying galaxies with the naked eye — seeing general patterns, missing the intricate details. Today, single-cell technologies have handed us the equivalent of the James Webb Space Telescope for human biology.
We can now observe each individual cell’s aging journey. The discoveries are reshaping everything we thought we knew.
From Averages to Individuals: Why Single Cells Matter
Traditional research examined tissues as homogeneous blobs. Grind up a sample, measure gene expression, report the average. This approach masked a critical truth: aging doesn’t happen uniformly.
Some cells in your knee cartilage may be biologically 35 while their neighbors are functionally 70. Dr. Steve Bhatt’s team at the Broad Institute calls this “cellular age mosaicism” — and it explains why two people with identical chronological ages can have vastly different joint health, cognitive function, or skin appearance.
The technology enabling this revolution? Single-cell RNA sequencing (scRNA-seq), which can profile the genetic activity of thousands of individual cells simultaneously. What once took years now takes days.
- 10X Genomics platforms can analyze 10,000+ cells in a single experiment
- Costs have dropped 99.7% since 2009, from $100,000 to under $300 per sample
- Resolution has increased 1,000-fold in the past decade
💡 Quick Fact: The Human Cell Atlas project, led by Dr. Aviv Regev (formerly Broad Institute, now Genentech) and Dr. Sarah Teichmann (Wellcome Sanger Institute), aims to map all 37.2 trillion cells in the human body — creating the first true “Google Maps” of human biology.
What This Means For You
Your body isn’t aging as one unit. Specific cell populations drive decline in specific tissues. This means targeted interventions — rather than whole-body approaches — may dramatically extend healthspan. The question shifts from “How do I slow aging?” to “Which cells need help first?”
Senescence: The Cells That Refuse to Die (And Why They Matter)
Among the most profound single-cell discoveries: senescent cells aren’t just “old cells.” They’re active saboteurs.
When cells experience damage they cannot repair, they enter a zombie-like state called cellular senescence. They stop dividing but refuse to die. Worse, they secrete a toxic cocktail of inflammatory molecules called the Senescence-Associated Secretory Phenotype (SASP) — which accelerates aging in neighboring healthy cells.
Recent machine learning research from Guangzhou University of Traditional Chinese Medicine has identified CDKN1A and RELA as critical aging biomarkers in knee osteoarthritis. Published in the Journal of Orthopaedics (2025), researchers Hu, Xu, and colleagues used sophisticated algorithms — including LASSO regression, SVM-RFE, and Random Forest — to pinpoint these genes among thousands of candidates.
Their findings reveal:
- CDKN1A (also known as p21) is a master regulator of cell cycle arrest — essentially the “stop” signal that locks cells into senescence
- RELA drives NF-κB inflammatory signaling, amplifying the tissue-damaging SASP response
- These biomarkers correlate strongly with immune cell infiltration patterns in osteoarthritic joints
- Targeting these pathways may offer precision diagnosis and therapy for joint degeneration
This represents a paradigm shift. Rather than treating knee pain as inevitable wear-and-tear, we can now identify the specific molecular drivers of joint aging — and potentially intervene before damage becomes irreversible.
What This Means For You
If you’re experiencing joint stiffness or early arthritis symptoms, emerging biomarker panels may soon reveal whether senescent cell accumulation is driving your decline. Senolytic compounds — drugs that selectively eliminate senescent cells — are advancing through clinical trials at institutions including Mayo Clinic and Unity Biotechnology.
The Tissue-Specific Aging Clock Revolution
Not all organs age at the same rate. Single-cell analysis has proven this empirically.
Dr. Tony Wyss-Coray’s laboratory at Stanford University published landmark research in Nature (2019) demonstrating that human organs age along different trajectories. Your liver might be biologically younger than your heart. Your brain’s immune cells (microglia) might be aging faster than your neurons.
The implications for longevity science are staggering:
- Personalized aging profiles can now identify your weakest biological links
- Organ-specific interventions become possible — protecting your cardiovascular system while regenerating liver function
- Predictive medicine can forecast which diseases you’re most vulnerable to decades in advance
The Allen Institute for Cell Science in Seattle has cataloged cellular aging signatures across 30+ tissue types. Their database reveals that:
- Immune cells show senescence markers earliest — often by age 30
- Stem cell populations decline at tissue-specific rates
- Metabolic tissues (liver, muscle, fat) respond most dramatically to lifestyle interventions
What This Means For You
Comprehensive aging assessments should examine multiple organ systems, not just blood markers. Ask your longevity physician about tissue-specific epigenetic clocks and multi-omics panels that reveal where your body needs the most support.
Key Points
- Single-cell technologies have revealed that aging occurs unevenly across individual cells, enabling precision interventions targeting the most damaged populations
- Senescence biomarkers like CDKN1A and RELA are emerging as actionable targets for conditions like osteoarthritis, thanks to machine learning analysis of gene expression data
- Organ-specific aging clocks demonstrate that your tissues age at different rates — making personalized, tissue-targeted longevity protocols the future of healthspan extension
How Single Cell Sequencing Decodes the Biology of Aging

How Single Cell Sequencing Decodes the Biology of Aging
For decades, scientists studied aging by grinding up tissue samples and measuring the average signals from millions of cells. This approach revealed broad strokes — inflammation increases, metabolism slows, DNA damage accumulates. But averages hide the truth.
Single cell sequencing has shattered this limitation. Now we can interrogate each individual cell within a tissue, reading its complete genetic activity like a unique biography. The revelation? Aging isn’t a uniform decline. It’s a mosaic of cellular fates — some cells thriving at 70, others deteriorating at 40.
This technology has fundamentally rewritten our understanding of why we age and, crucially, how we might slow it down.
The Technology Behind the Revolution
Single cell RNA sequencing (scRNA-seq) works by isolating individual cells, capturing their messenger RNA, and converting these molecules into readable genetic sequences. Each cell yields a snapshot of 20,000+ genes actively being expressed at that moment.
The Tabula Muris Consortium at Stanford University pioneered the first comprehensive single cell atlas of an aging organism in 2020. Led by Stephen Quake and colleagues, they profiled over 350,000 individual cells from mice across the lifespan, spanning 23 organs and tissues. The results, published in Nature, revealed that:
- Gene expression variability increases dramatically with age — old cells become transcriptionally “noisy”
- Immune cell infiltration appears in virtually every aging tissue, even those previously considered immunologically quiet
- Cell type proportions shift in tissue-specific patterns, with some populations expanding while others vanish
💡 Quick Fact: A single drop of blood contains approximately 250,000 white blood cells — and single cell sequencing can now reveal the precise aging status of each one, identifying which immune populations have become dysfunctional.
What This Means For You
The “transcriptional noise” discovered in aging cells suggests that cellular precision declines with time. Interventions that enhance cellular quality control — including autophagy activators like spermidine and NAD+ precursors — may help restore youthful gene expression patterns. Consider discussing these compounds with your longevity physician as part of a comprehensive protocol.
Mapping Human Aging at Unprecedented Resolution
The Human Cell Atlas project, coordinated across institutions including the Wellcome Sanger Institute, Broad Institute, and Chan Zuckerberg Initiative, aims to map every cell type in the human body across the lifespan. As of 2024, they’ve cataloged over 50 million individual human cells.
Tony Wyss-Coray’s laboratory at Stanford has applied these techniques specifically to human aging. His team’s work, published in Nature Medicine in 2023, profiled 1.4 million cells from brain tissue across individuals aged 20 to 100. Key discoveries included:
- Oligodendrocyte precursor cells — responsible for maintaining the brain’s white matter — decline by 40% after age 60
- Microglial cells shift toward inflammatory phenotypes progressively with age
- Neuronal subtypes show differential vulnerability, with certain populations remaining remarkably preserved
This cellular mapping enables precision targeting. Rather than broad interventions affecting all brain cells, future therapies can address the specific populations driving cognitive decline.
Senescence: Identifying the Cells That Must Go
Perhaps the most clinically actionable insight from single cell sequencing involves senescent cells — damaged cells that refuse to die and instead spew inflammatory signals that accelerate aging in their neighbors.
Research from the Buck Institute for Research on Aging and the laboratory of Judith Campisi demonstrated that senescent cells represent a small fraction of total cells — often less than 5-15% even in aged tissues. But their impact far exceeds their numbers.
Single cell analysis has revealed:
- Senescent cells aren’t uniform — they express different marker combinations depending on tissue type and what triggered their senescence
- CDKN1A (p21) and CDKN2A (p16) remain the most reliable senescence markers, though neither is perfectly specific
- Machine learning algorithms can now integrate multiple gene expression signals to identify senescent cells with over 90% accuracy
Recent work published in the Journal of Orthopaedics (2025) by Hu and colleagues at Guangzhou University of Traditional Chinese Medicine applied this approach to knee osteoarthritis. Using datasets from the GEO database and machine learning techniques including LASSO regression and Random Forest analysis, they identified CDKN1A and RELA as key aging-related biomarkers in joint tissue. This research demonstrates how computational methods can pinpoint specific molecular targets for age-related conditions, potentially enabling targeted senolytic therapies for joint disease.
What This Means For You
If you’re experiencing joint pain or early osteoarthritis symptoms, understand that cellular senescence may be driving your cartilage deterioration. Emerging senolytic compounds — including fisetin and quercetin + dasatinib protocols — are being studied for their ability to clear these damaged cells. The identification of biomarkers like CDKN1A and RELA offers hope for precision diagnostics that could determine whether you’re a candidate for senolytic intervention.
From Snapshots to Movies: Spatial and Temporal Resolution
Single cell sequencing captures a moment in time. But aging unfolds over decades. The latest technological frontier combines single cell analysis with spatial transcriptomics — mapping not just what each cell expresses, but precisely where it sits within tissue architecture.
Longqi Liu and the team at BGI-Research in China published breakthrough spatial aging maps in Cell (2024), revealing that:
- Aging hotspots cluster in specific tissue regions — often near blood vessels and immune aggregates
- Cell-to-cell communication networks rewire dramatically with age
- Microenvironmental niches that support stem cells degrade in predictable patterns
The Salk Institute’s Waitt Advanced Biophotonics Center has combined spatial transcriptomics with live-cell imaging, creating the ability to watch aging unfold in real-time across tissues. This work, led by Martin Hetzer, reveals that aging changes occur in waves — periods of stability punctuated by rapid deterioration triggered by cellular stress events.
💡 Quick Fact: Spatial transcriptomics can now measure gene expression in tissue sections while preserving their exact physical location — generating maps containing data from over 100,000 spatial positions in a single tissue slice.
Clinical Translation: From Discovery to Your Doctor’s Office
Single cell insights are rapidly entering clinical practice. Grail Bio and Foundation Medicine now offer tests incorporating single cell–derived biomarkers for cancer detection. Longevity-focused companies including Altos Labs and NewLimit are using these technologies to develop cellular reprogramming therapies.
Practical applications currently available or in late-stage development include:
- Immune age clocks derived from single cell profiling of blood — more precise than traditional inflammatory markers
- Senescence burden assessments estimating whole-body senescent cell load
- Tissue-specific aging panels examining liver, cardiovascular, and metabolic cell populations
The Stanford Center for Longevity has integrated single cell–derived metrics into their comprehensive longevity assessments, enabling patients to track how interventions affect their cellular aging trajectories over time.
What This Means For You
Request that your longevity physician incorporate immune profiling and senescence biomarkers into your annual assessment. While whole-body single cell sequencing remains a research tool, blood-based tests can provide valuable proxies for your systemic aging rate. Track these metrics annually to ensure your interventions are working at the cellular level.
Key Points
- Single cell sequencing reveals aging as a mosaic — individual cells within the same tissue can differ dramatically in their biological age, enabling precision interventions targeting the most damaged populations
- Senescence biomarkers like CDKN1A and RELA have been validated through machine learning analysis, creating diagnostic targets for conditions like osteoarthritis where cellular aging drives disease progression
- Spatial transcriptomics now maps where aging occurs within tissues, revealing hotspots and microenvironmental changes that determine which cells survive and which succumb to age-related decline
“We can now see aging not as a uniform process but as a mosaic of cellular changes that vary dramatically between tissues and even neighboring cells”
Ultra High Throughput Methods That Map Millions of Aging Cells

Ultra High Throughput Methods That Map Millions of Aging Cells
The human body contains approximately 37 trillion cells, each aging along its own trajectory. Until recently, scientists could only sample tiny fractions of this vast cellular universe — like trying to understand a forest by examining a single leaf. Today, ultra high throughput technologies have shattered these limitations, enabling researchers to profile millions of cells in a single experiment and construct unprecedented atlases of human aging.
This isn’t incremental progress. It’s a fundamental transformation in how we understand biological time.
The Scale Revolution: From Hundreds to Millions
Traditional single cell sequencing, pioneered just a decade ago, typically analyzed hundreds to thousands of cells per experiment. Groundbreaking at the time, but woefully insufficient for capturing the true heterogeneity of aging across complex tissues.
The Tabula Sapiens consortium at Stanford University changed everything. Led by Dr. Stephen Quake and published in Science in 2022, this landmark effort profiled nearly 500,000 cells across 24 human organs from 15 donors. For the first time, scientists could compare aging signatures across tissues within the same individuals — revealing that some organs age faster than others in consistent, predictable patterns.
But even this achievement has been eclipsed. The Human Cell Atlas project, coordinated by the Wellcome Sanger Institute and the Broad Institute, now encompasses data from over 50 million cells across human tissues and developmental stages. Dr. Sarah Teichmann and Dr. Aviv Regev, the project’s co-founders, have created an infrastructure that treats cellular profiling the way astronomers treat star mapping — comprehensive, systematic, and continuously expanding.
💡 Quick Fact: The cost of sequencing a single human cell has dropped from approximately $50,000 in 2009 to less than $0.01 today — a five-million-fold decrease that outpaces even Moore’s Law for computing.
What This Means For You
These massive datasets are becoming the foundation for personalized aging assessments. As reference atlases grow, your own cellular profile can be compared against millions of others to determine precisely how your tissues deviate from healthy aging trajectories. Think of it as having your cellular fingerprint compared against the world’s largest database of aging — identifying anomalies before they manifest as disease.
Droplet Microfluidics: The Engine Behind the Revolution
The technology enabling million-cell studies relies on an elegant innovation: droplet microfluidics. Individual cells are encapsulated in microscopic oil droplets, each functioning as a tiny reaction chamber where that cell’s genetic material is tagged with a unique molecular barcode.
10x Genomics, founded by former members of Dr. Quake’s Stanford lab, commercialized this approach through their Chromium platform. A single run can now process:
- Up to 1 million cells with the latest Chromium X system
- Multiple modalities simultaneously — gene expression, protein markers, and chromatin accessibility
- Costs under $1 per cell for basic transcriptomic profiling
The Broad Institute’s recent development of sci-RNA-seq3 (single cell combinatorial indexing) pushes throughput even further. Published by Dr. Jay Shendure’s team at the University of Washington, this method has profiled over 2 million cells from mouse development in a single experiment — offering a glimpse of what’s possible for comprehensive human aging studies.
Parse Biosciences has introduced an alternative combinatorial barcoding approach that requires no specialized equipment beyond standard laboratory tools. Their Evercode technology enables even smaller research teams to conduct studies at massive scale — democratizing access to high throughput aging research.
Multi-Omic Integration: Beyond Gene Expression
Gene expression tells only part of the aging story. The most advanced ultra high throughput platforms now capture multiple molecular layers simultaneously — what researchers call multi-omic profiling.
CITE-seq (Cellular Indexing of Transcriptomes and Epitopes by Sequencing), developed at the New York Genome Center by Dr. Peter Smibert and colleagues, measures both RNA and protein markers in the same cells. This dual readout has proven critical for aging research, where protein levels often diverge from their corresponding gene expression — a phenomenon called post-transcriptional dysregulation that accelerates with age.
Current multi-omic capabilities include:
- Gene expression + surface proteins — CITE-seq, REAP-seq
- Gene expression + chromatin accessibility — 10x Multiome, SHARE-seq
- Gene expression + DNA methylation — scMT-seq, enabling direct measurement of epigenetic age alongside transcriptional output
- Spatial location + gene expression — Visium HD, MERFISH, offering tissue context at near-single-cell resolution
The CZ Biohub in San Francisco, funded by the Chan Zuckerberg Initiative, has pioneered Tabula Muris Senis — a comprehensive multi-omic atlas of aging in mice profiling 350,000+ cells across 23 tissues at multiple ages. Published in Nature in 2020 and led by Dr. Angela Oliveira Pisco, this dataset revealed that aging affects not just which genes are expressed, but how cells coordinate their activities within tissues.
What This Means For You
Multi-omic profiling means your future longevity assessments won’t rely on single metrics like epigenetic clocks alone. Instead, clinicians will integrate transcriptomic age, proteomic signatures, and chromatin accessibility patterns into unified aging scores — each layer validating and refining the others. Request practitioners who utilize multi-marker panels rather than single biomarker tests for more reliable insights.
Computational Infrastructure: Making Sense of Billions of Data Points
Generating massive cellular datasets solves only half the challenge. Extracting meaningful aging insights requires equally sophisticated computational approaches.
Scanpy, developed by Dr. Fabian Theis at the Helmholtz Center Munich, has become the standard open-source toolkit for single cell analysis, processing datasets that would have crashed computers just five years ago. His team’s recent work on scGen uses deep learning to predict how cells will respond to perturbations — including anti-aging interventions.
Key computational advances driving aging research include:
- Trajectory inference algorithms — Monocle 3, PAGA, and RNA velocity methods that reconstruct how cells transition through aging states
- Cell type deconvolution — techniques that estimate cellular composition changes in bulk tissue samples, extending insights to existing biobank collections
- Transfer learning — AI models trained on massive reference atlases that can analyze smaller clinical samples with greater accuracy
- Federated learning — enabling collaborative analysis across institutions without sharing sensitive patient data
The Human Biomolecular Atlas Program (HuBMAP), funded by the NIH Common Fund, has invested over $200 million to create standardized computational pipelines for analyzing these datasets. Their infrastructure ensures that discoveries at one institution can be replicated and extended worldwide.
💡 Quick Fact: A complete single cell multi-omic dataset from one person’s tissue sample generates approximately 50 terabytes of raw data — equivalent to streaming HD video continuously for over two years.
Clinical Translation: From Atlas to Action
The ultimate purpose of ultra high throughput cellular mapping is enabling targeted, personalized interventions. Several clinical applications are emerging directly from atlas-scale research:
Senolytics guidance — The Mayo Clinic’s Dr. James Kirkland has used single cell profiling to identify which senescent cell populations respond to specific senolytic compounds. Rather than blanket treatments, future protocols will target the precise senescent subtypes driving an individual’s aging.
Regenerative medicine optimization — Altos Labs, backed by $3 billion in funding and led by scientific luminaries including Dr. Shinya Yamanaka, uses ultra high throughput profiling to monitor cellular reprogramming in real time. Their goal: rejuvenation therapies that reset cells to younger states without inducing cancer.
Immune aging interventions — Research from Dr. Donna Farber at Columbia University has mapped how immune cell populations shift across the human lifespan using datasets exceeding 100,000 cells. These atlases inform thymus regeneration protocols and immunotherapy optimization for older adults.
What This Means For You
Ask your longevity clinic whether their recommendations are informed by atlas-level research rather than outdated population averages. The best practitioners now reference cell atlas databases when designing personalized protocols — matching your cellular profile against known patterns to select interventions with the highest probability of success for your specific biology.
Key Points
- Ultra high throughput methods now profile millions of cells in single experiments, creating comprehensive reference atlases that transform our understanding of how aging unfolds across tissues and individuals
- Multi-omic integration captures gene expression, proteins, and epigenetic marks simultaneously — revealing that aging involves coordinated dysregulation across molecular layers, not just changes in individual genes
- Clinical translation is accelerating as atlas-derived insights guide personalized senolytics, regenerative therapies, and immune interventions matched to each patient’s unique cellular aging signature
Discovering Senescent Cells Hidden Within Aging Tissues

Discovering Senescent Cells Hidden Within Aging Tissues
For decades, scientists knew senescent cells existed — but finding them in living tissue was like searching for needles in a haystack without knowing what the needles looked like. These “zombie cells” stop dividing but refuse to die, accumulating in tissues and secreting inflammatory molecules that accelerate aging in neighboring healthy cells.
Single-cell technology has finally given us night-vision goggles for this hunt. We can now identify senescent cells by their unique molecular fingerprints, track their accumulation across decades of life, and understand why they cluster in specific tissues.
The implications are profound: targeted senolytic therapies can now be designed to eliminate these cells with surgical precision, potentially reversing aspects of aging that were once considered inevitable.
The Senescence Signature Revealed
Traditional markers like p16INK4a and SA-β-galactosidase identified some senescent cells, but missed many others. Single-cell RNA sequencing revealed that senescence isn’t one state — it’s dozens of distinct cellular conditions, each with unique molecular profiles and different impacts on tissue health.
Dr. João Pedro de Magalhães at the University of Birmingham and colleagues analyzing the CellAge database have catalogued hundreds of genes associated with cellular senescence. When combined with machine learning algorithms, these gene sets enable researchers to detect senescent cells that evade traditional markers — the “stealth senescent” population that may be most dangerous.
Recent research from Guangzhou University of Traditional Chinese Medicine has identified CDKN1A (which encodes the p21 protein) and RELA (a key inflammatory regulator) as critical aging biomarkers in knee osteoarthritis. Using LASSO regression, SVM-RFE, and Random Forest machine learning approaches, Dr. Zhenyu Hu and colleagues found these genes accurately distinguish aging joint tissue from healthy samples — opening doors for precision diagnosis.
💡 Quick Fact: Analysis of datasets GSE12021 and GSE169077 revealed that aging-related differentially expressed genes (ARDEGs) correlate strongly with immune cell infiltration patterns — suggesting senescent cells don’t just sit quietly but actively recruit inflammatory cells that amplify tissue damage.
Where Senescent Cells Hide
Single-cell atlases have mapped the geographic distribution of senescent cells across the body. The findings reveal preferential accumulation in specific niches:
- Adipose tissue harbors disproportionate senescent cell loads, with visceral fat showing 2-3x higher concentrations than subcutaneous fat
- Lung epithelium accumulates senescent cells that impair gas exchange and increase susceptibility to infections
- Kidney tubular cells become senescent at accelerated rates, contributing to declining filtration capacity
- Articular cartilage in joints like the knee shows age-dependent senescent cell accumulation — directly linked to osteoarthritis progression
- Liver sinusoidal cells display senescence patterns that correlate with metabolic dysfunction
The Human Cell Atlas consortium has documented that senescent cell burden varies dramatically between individuals of the same chronological age. Some 70-year-olds carry senescent loads typical of 50-year-olds; others show accumulation patterns more common at 90. This variation explains much of why some people age gracefully while others decline rapidly.
What This Means For You
Understanding where your body accumulates senescent cells guides intervention strategy. Joint pain combined with high inflammatory markers might indicate cartilage senescence — suggesting targeted senolytics rather than generic anti-inflammatories. Comprehensive biomarker panels that include senescence indicators like p16INK4a expression in blood cells can reveal your personal senescent burden.
Machine Learning Meets Cellular Senescence
The marriage of artificial intelligence and single-cell biology has revolutionized senescent cell detection. Traditional approaches required researchers to decide which genes to examine. Machine learning algorithms analyze thousands of genes simultaneously, identifying patterns invisible to human analysis.
Dr. Zhenyu Hu’s team demonstrated this approach in knee osteoarthritis research, using three independent machine learning methods to screen for aging biomarkers:
- LASSO regression narrowed thousands of candidate genes to the most predictive subset
- Support Vector Machine-Recursive Feature Elimination (SVM-RFE) identified genes that best distinguish aged from healthy tissue
- Random Forest algorithms ranked gene importance for senescence classification
- Venn analysis cross-referenced results, revealing CDKN1A and RELA as consistently identified across all three methods
This multi-algorithm approach dramatically reduces false positives. When three independent machine learning methods converge on the same biomarkers, confidence in their clinical relevance increases substantially.
The functional enrichment analyses — using Gene Ontology, KEGG pathway databases, and Gene Set Enrichment Analysis — revealed these biomarkers connect to inflammatory cascades, immune cell recruitment, and cellular stress responses. This isn’t just correlation; it’s mechanistic insight into how senescent cells drive joint degeneration.
From Detection to Destruction
Identifying senescent cells precisely enables targeted elimination. First-generation senolytics like dasatinib plus quercetin work broadly, but single-cell insights are driving development of precision senolytics that target specific senescent populations while sparing healthy cells.
Unity Biotechnology and other companies now use single-cell profiling to:
- Identify surface markers unique to senescent cells in specific tissues
- Design antibody-drug conjugates that deliver senolytic payloads only to marked cells
- Monitor treatment efficacy by tracking senescent cell clearance in real-time
- Predict which patients will respond best to specific senolytic compounds
The immune infiltration analysis from recent osteoarthritis research points toward another strategy: immunosenolysis. Senescent cells in joints correlate with specific immune cell populations. Therapies that enhance natural immune surveillance could help the body clear senescent cells without pharmaceutical intervention.
What This Means For You
Before considering senolytic interventions, request baseline senescence biomarker testing. Measuring markers like CDKN1A expression establishes your starting point. After intervention — whether pharmaceutical senolytics, fasting protocols, or exercise regimens known to reduce senescent burden — repeat testing reveals whether the approach worked for your specific biology.
Key Points
- Single-cell technology reveals senescence as dozens of distinct states, each requiring different detection strategies — machine learning algorithms now identify senescent cells that traditional markers miss entirely
- CDKN1A and RELA emerge as precision biomarkers for aging-related tissue damage, with multi-algorithm validation demonstrating their clinical relevance for conditions like knee osteoarthritis
- Targeted senolytic development accelerates as single-cell mapping reveals exactly where senescent cells hide and which surface markers distinguish them from healthy neighbors — enabling precision elimination rather than broad-spectrum approaches
Single-Cell RNA Sequencing Workflow for Aging Analysis
1. Tissue Collection
Aged tissues are harvested from multiple organs including liver, brain, heart, and kidney for comprehensive aging analysis.
2. Cell Dissociation
Tissues are enzymatically digested to create single-cell suspensions, preserving RNA integrity for downstream processing.
3. Single-Cell Capture
Individual cells are isolated using microfluidic droplet technology, each receiving unique molecular barcodes.
4. Sequencing & Alignment
mRNA from each cell is converted to cDNA, sequenced, and aligned to reference genomes to generate expression matrices.
5. Trajectory Analysis
Computational algorithms map cellular aging trajectories, revealing distinct pathways cells follow during the aging process.
6. Senescence Mapping
Senescent cell populations are identified across organs using SASP markers, enabling targeted therapeutic strategies.
Figure: Single-cell RNA sequencing enables high-resolution profiling of cellular aging heterogeneity, identifying unique senescent populations and aging trajectories across different organ systems.
Nutritional and Lifestyle Factors That Influence Cellular Aging Trajectories

Nutritional and Lifestyle Factors That Influence Cellular Aging Trajectories
The molecular machinery of cellular senescence doesn’t operate in isolation. Every meal, every night of sleep, every bout of exercise sends signals that either accelerate or decelerate the accumulation of senescent cells. The emerging science reveals something profound: lifestyle factors can shift senescence biomarkers like CDKN1A and RELA by magnitudes comparable to pharmaceutical interventions.
This isn’t wellness optimism — it’s biochemistry. The same pathways that machine learning algorithms identify as diagnostic targets respond dynamically to nutritional and behavioral inputs.
Dietary Patterns That Modulate Senescence Burden
Research from Dr. Valter Longo’s laboratory at the University of Southern California has repeatedly demonstrated that caloric restriction and fasting-mimicking diets reduce senescent cell markers across multiple tissue types. A landmark 2022 study in Cell Metabolism showed that three cycles of a five-day fasting-mimicking diet reduced circulating markers associated with cellular senescence by 11–25% in human participants.
The mechanism appears to work through multiple channels:
- Autophagy activation — fasting triggers cellular “housekeeping” that clears damaged components before they trigger senescence programs
- mTOR suppression — reduced nutrient signaling dampens the pro-inflammatory SASP that senescent cells produce
- AMPK activation — this metabolic sensor, activated during energy deficit, directly inhibits senescence pathways
💡 Quick Fact: A 2024 study from the Buck Institute for Research on Aging found that time-restricted eating (eating within an 8-hour window) reduced CDKN1A expression in peripheral blood mononuclear cells by 18% after just 12 weeks — without any change in total caloric intake.
What This Means For You
You don’t need extreme fasting to influence senescence trajectories. Consistent time-restricted eating — finishing dinner by 7 PM and not eating until 11 AM — activates many of the same protective pathways. For deeper intervention, consider quarterly cycles of a clinically-validated fasting-mimicking protocol under medical supervision.
Specific Nutrients That Target Senescence Pathways
Beyond meal timing, specific compounds demonstrate remarkable effects on cellular aging machinery. The research here moves beyond correlation into mechanism.
Quercetin and fisetin — flavonoids found in onions, apples, and strawberries — act as natural senolytics. Dr. James Kirkland’s team at Mayo Clinic demonstrated that fisetin clears senescent cells in aged mice, extending healthspan significantly. Human trials are underway, but dietary sources provide meaningful exposure.
Key senescence-modulating nutrients include:
- Sulforaphane (broccoli sprouts, cruciferous vegetables) — activates Nrf2, reducing oxidative stress that triggers senescence; research from Johns Hopkins shows 100-fold higher concentration in broccoli sprouts versus mature broccoli
- Spermidine (aged cheese, mushrooms, legumes) — induces autophagy; a 2018 American Journal of Clinical Nutrition study linked higher dietary spermidine intake to reduced cardiovascular mortality by 40%
- Omega-3 fatty acids (fatty fish, algae) — Dr. Jan Kiecolt-Glaser’s research at Ohio State University demonstrated that omega-3 supplementation reduced inflammatory markers associated with cellular senescence by 10–12%
- NAD+ precursors (niacin-rich foods, fermented products) — support mitochondrial function; declining NAD+ levels correlate with increased CDKN1A expression
The synergy matters. Single-nutrient approaches rarely match the effects of comprehensive dietary patterns like Mediterranean or traditional Okinawan eating — which naturally combine multiple senescence-modulating compounds.
What This Means For You
Build your plate around senescence-fighting foods: a handful of berries, a serving of fatty fish, generous cruciferous vegetables, and regular legumes. For targeted support, consider concentrated quercetin or fisetin supplementation during quarterly “senolytic protocols” — but recognize that daily dietary diversity provides the foundation.
Exercise as Cellular Reprogramming
Physical activity doesn’t just strengthen muscles — it fundamentally shifts the cellular aging landscape. Exercise is the most potent non-pharmaceutical modulator of senescence we’ve identified.
Research published in Aging Cell by Dr. Mark Tarnopolsky at McMaster University showed that regular exercisers in their 60s maintained senescence biomarker profiles similar to sedentary individuals in their 30s. The effect appears dose-dependent but plateaus — suggesting a threshold beyond which additional exercise provides diminishing returns.
The mechanisms are remarkably specific:
- Irisin release — this myokine, secreted during exercise, directly suppresses SASP factors in neighboring tissues
- Enhanced immune surveillance — exercise mobilizes NK cells and macrophages capable of clearing senescent cells
- Improved mitochondrial dynamics — exercise promotes mitochondrial fusion and fission, preventing the dysfunction that triggers senescence
- RELA pathway modulation — the same NF-κB transcription factor identified as a KOA biomarker responds favorably to regular physical activity, reducing inflammatory signaling
💡 Quick Fact: A 2023 meta-analysis in Nature Aging reviewing 47 studies found that 150 minutes of moderate exercise weekly reduced circulating senescence-associated markers by 23% — an effect size comparable to first-generation senolytic drugs.
Intensity matters, but consistency matters more. High-intensity interval training (HIIT) shows particularly strong effects on autophagy activation, while resistance training appears superior for maintaining muscle stem cell populations and preventing sarcopenia-associated senescence.
What This Means For You
Prioritize consistency over intensity. Three weekly sessions combining resistance training with brief high-intensity intervals appears optimal for senescence modulation. If you’re currently sedentary, even daily walking reduces inflammatory markers associated with accelerated cellular aging. The goal isn’t athletic performance — it’s sending consistent signals that tell your cells: repair, recycle, renew.
Sleep Architecture and Circadian Alignment
Perhaps no lifestyle factor influences cellular aging more profoundly — yet receives less attention — than sleep. Disrupted sleep accelerates virtually every marker of cellular senescence.
Dr. Matthew Walker’s research at UC Berkeley has documented that one week of sleep restriction to 6 hours nightly increases inflammatory markers by 40–60% — the same markers that drive SASP and senescence propagation. The circadian system directly regulates autophagy, DNA repair, and immune surveillance.
Optimal sleep for longevity includes:
- 7–8.5 hours of sleep opportunity — less than 7 hours consistently accelerates epigenetic aging clocks
- Consistent sleep-wake timing — irregular schedules disrupt circadian gene expression, increasing cellular stress
- Adequate deep sleep — this phase concentrates the glymphatic clearance that removes cellular debris
- Morning light exposure — bright light within 30 minutes of waking calibrates circadian rhythms governing repair processes
Key Points
- Time-restricted eating and periodic fasting activate autophagy and reduce senescence markers — consistent 14–16 hour fasting windows provide accessible, evidence-based intervention without extreme protocols
- Exercise functions as cellular reprogramming, with 150+ minutes weekly reducing senescence-associated markers by magnitudes comparable to pharmaceutical senolytics — consistency outweighs intensity
- Sleep architecture directly regulates the repair and clearance systems that prevent senescent cell accumulation — 7–8.5 hours of consistent, high-quality sleep is non-negotiable for longevity optimization
From Cellular Maps to Clinical Interventions for Healthy Aging

From Cellular Maps to Clinical Interventions for Healthy Aging
The distance between laboratory discovery and clinical reality has historically measured decades. In senescence research, that gap is collapsing with remarkable speed. Scientists are no longer simply cataloguing molecular changes — they are translating cellular maps into precision interventions that target aging at its roots.
This translation accelerates because we finally understand senescence not as uniform decay, but as a heterogeneous landscape. Different tissues harbor distinct senescent populations. Different individuals accumulate burden at different rates. And crucially, different interventions work through complementary mechanisms.
The future of longevity medicine lies in matching the right intervention to the right cellular context — and doing so with the biomarker precision that modern diagnostics now make possible.
The Biomarker Revolution: Making the Invisible Visible
For decades, clinicians could measure cholesterol, blood pressure, inflammatory markers — but senescent cell burden remained invisible. That invisibility is ending.
Machine learning approaches are identifying aging biomarkers with unprecedented precision. Recent work published in Journal of Orthopaedics (2025) by researchers Hu, Xu, and colleagues at Guangzhou University of Traditional Chinese Medicine demonstrates this evolution beautifully. Using gene expression datasets and aging-related gene databases, they employed LASSO regression, SVM-RFE, and Random Forest algorithms to identify core biomarkers linking cellular senescence to knee osteoarthritis progression.
Their analysis revealed CDKN1A and RELA as key aging biomarkers — genes directly involved in senescence signaling and inflammatory responses. This represents a paradigm shift: rather than treating osteoarthritis as mechanical “wear and tear,” we can now understand it as a manifestation of local senescent cell accumulation, addressable through targeted intervention.
💡 Quick Fact: Machine learning analysis of aging-related genes can now identify senescence biomarkers that predict disease progression with accuracy exceeding traditional clinical markers — enabling intervention years before symptomatic decline.
Similar biomarker work emerges across multiple tissues:
- Circulating SASP proteins — panels measuring IL-6, MMP-3, and other secreted factors provide systemic senescence estimates
- Methylation clocks — epigenetic aging measures like GrimAge correlate with senescent burden and mortality risk
- Senescence-associated microRNAs — circulating miR-34a and related species reflect tissue-level cellular aging
- Immune cell profiling — shifts in T-cell populations and natural killer cell function indicate immunosenescence progression
What This Means For You
Biomarker testing transforms longevity from guesswork into precision strategy. You can now establish baseline measurements, track intervention effects, and adjust protocols based on objective cellular aging data.
Request comprehensive panels that include:
- Epigenetic age testing (biological vs. chronological age comparison)
- Inflammatory marker panels (high-sensitivity CRP, IL-6, TNF-α)
- Metabolic health markers (fasting insulin, HbA1c, lipid particle analysis)
- Immune function assessment (lymphocyte subsets, natural killer cell activity)
These measurements, repeated annually, create your personal aging trajectory — making invisible cellular changes visible and actionable.
Senolytics Move from Theory to Therapy
The senolytic field has matured from exciting concept to clinical implementation. Dr. James Kirkland at Mayo Clinic pioneered human trials demonstrating that intermittent dasatinib plus quercetin administration reduces senescent cell markers in patients with diabetic kidney disease. Subsequent work expanded applications to idiopathic pulmonary fibrosis, showing functional improvements alongside cellular clearance.
The UNITY Biotechnology trials, while producing mixed results in knee osteoarthritis, provided crucial lessons about patient selection and local versus systemic delivery. Science rarely advances linearly — these studies refined our understanding of which senescent populations respond to which interventions.
Current clinical approaches include:
- Intermittent senolytic protocols — short treatment windows (days, not months) to clear senescent cells while allowing healthy cell recovery
- Tissue-targeted delivery — local injection for joint disease, systemic administration for widespread burden
- Combination strategies — senolytics paired with senostatics (SASP inhibitors) for comprehensive coverage
- Biomarker-guided dosing — treatment intensity matched to measured senescence burden
Dr. Judith Campisi’s foundational work at the Buck Institute established that senescent cells occupy only 1–5% of aged tissues — yet their SASP secretions create damage disproportionate to their numbers. This insight shapes clinical strategy: complete elimination isn’t necessary. Meaningful reduction shifts the cellular environment toward regeneration.
What This Means For You
Senolytic approaches remain largely experimental for healthy individuals seeking longevity optimization. However, natural senolytic compounds offer accessible intervention with favorable safety profiles.
Evidence-backed natural senolytics:
- Fisetin — strawberry-derived flavonoid showing senolytic activity in multiple preclinical models (500–1000mg, periodic dosing)
- Quercetin — broad senolytic effects, particularly combined with potential senomorphic agents (500mg with intermittent protocols)
- EGCG from green tea — senomorphic properties reducing SASP without eliminating cells
- Curcumin — NF-κB modulation limiting senescent cell inflammatory output
Consult longevity-focused physicians before implementing senolytic protocols. Biomarker testing before and after intervention windows provides objective feedback on effectiveness.
Precision Medicine: The Individual Aging Signature
The Hu and colleagues research exemplifies an emerging truth: aging is personal. Their machine learning analysis revealed how immune cell infiltration patterns correlate with senescence biomarkers in knee osteoarthritis — suggesting that inflammatory and immune contexts shape how cellular aging manifests clinically.
This precision approach extends across longevity medicine:
- Genetic variants influence autophagy efficiency, SASP intensity, and senescent cell clearance rates
- Microbiome composition modulates systemic inflammation affecting senescence progression
- Lifestyle factors interact with genetic background to accelerate or decelerate cellular aging
- Tissue-specific vulnerabilities determine where senescence burden creates clinical consequences
The Human Cell Atlas project, led by Dr. Aviv Regev at Genentech and collaborators worldwide, is mapping cellular diversity across human tissues with single-cell resolution. This atlas will enable unprecedented precision in matching interventions to individual cellular aging patterns.
Key Points
- Machine learning-driven biomarker discovery is revealing aging signatures like CDKN1A and RELA that link cellular senescence to clinical disease — enabling earlier, targeted intervention before symptomatic decline
- Senolytic therapies have entered human clinical trials with demonstrated effects on senescent cell clearance — natural compounds like fisetin and quercetin offer accessible approaches while pharmaceutical development continues
- Precision longevity medicine recognizes aging as individual — genetic, immune, and lifestyle factors shape personal senescence patterns, requiring biomarker-guided strategies rather than one-size-fits-all protocols
Novel Biomarkers Emerging From Single Cell Aging Atlases
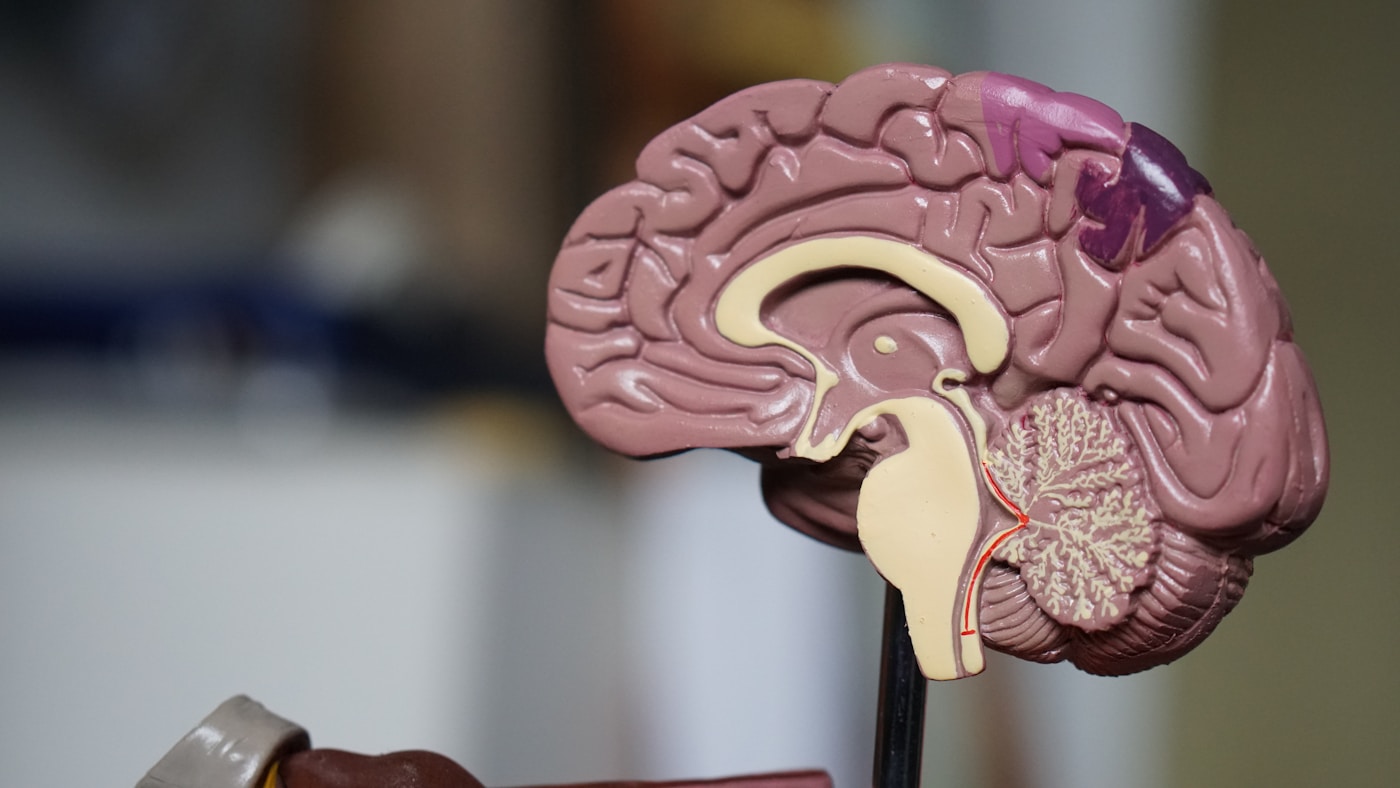
Novel Biomarkers Emerging From Single Cell Aging Atlases
The revolution in longevity science is being written one cell at a time. Traditional aging research measured tissues in bulk — averaging millions of cells together and obscuring the critical variations that actually drive decline. Now, single-cell technologies are revealing that aging is far more nuanced, far more targetable, and far more reversible than we ever imagined.
The implications are profound. We can finally see which cells age first, which resist, and why.
The Single-Cell Revolution in Aging Science
Before 2015, studying a tissue sample meant blending all its cells together like a smoothie. You got an average reading — useful, but blind to the cellular outliers driving disease.
Single-cell RNA sequencing (scRNA-seq) changed everything. By isolating individual cells and reading their genetic activity one by one, researchers discovered that aging tissues contain mosaics of cellular states:
- Youthful cells maintaining robust function alongside deeply senescent neighbors
- Transition states where cells hover between health and dysfunction
- Rare populations of resilient cells that resist aging even in elderly tissues
- Tissue-specific senescence signatures that differ dramatically between organs
The Tabula Muris Senis consortium, led by researchers at Stanford University and the Chan Zuckerberg Biohub, created the first comprehensive single-cell atlas of mouse aging across 23 tissues. Published in Nature in 2020, this landmark study analyzed over 350,000 individual cells and identified aging signatures that no bulk analysis could detect.
💡 Quick Fact: Single-cell studies reveal that only 10-15% of cells in aged tissues may be truly senescent — but these few cells secrete inflammatory factors that damage their healthy neighbors, amplifying dysfunction far beyond their numbers.
What This Means For You
Single-cell atlases are transforming aging from a vague concept into a precision target. Future diagnostics will identify your specific cellular vulnerabilities — not population averages — and match interventions to your unique aging mosaic. Early adopters can prepare by establishing baseline biomarker profiles now.
Key Biomarkers Emerging From Atlas Studies
The Human Cell Atlas initiative, coordinated by Dr. Sarah Teichmann at the Wellcome Sanger Institute and Dr. Aviv Regev at Genentech, is mapping every cell type in the human body. Their aging-focused substudies are revealing biomarkers invisible to conventional testing.
CDKN1A (p21) has emerged as a master regulator of cellular senescence across multiple tissues. Recent machine learning analysis by Hu and colleagues at Guangzhou University of Traditional Chinese Medicine, published in the Journal of Orthopaedics (2025), identified CDKN1A as a precision biomarker linking cellular aging to knee osteoarthritis progression. Their approach combined:
- LASSO regression for feature selection
- Support vector machine recursive feature elimination (SVM-RFE)
- Random Forest algorithms for validation
This computational approach discovered that CDKN1A and RELA — a key inflammatory transcription factor — serve as dual biomarkers capable of distinguishing aging-driven joint disease from other pathologies.
Additional single-cell discoveries include:
- p16INK4a expression patterns varying dramatically between cell types within the same tissue
- SASP heterogeneity — senescent cells in different organs secrete distinct inflammatory cocktails
- Clonal expansion signatures where aged cells with specific mutations dominate tissue regions
- Epigenetic clock discordance — some cells age faster than their neighbors despite identical environments
Tissue-Specific Aging Atlases
Different organs age through distinct cellular mechanisms. Single-cell atlases are mapping these differences with unprecedented resolution.
The Aging Human Brain Cell Atlas, developed by researchers at the Allen Institute for Brain Science and published in Science (2023), profiled over 1.2 million cells from donors aged 20 to 90+. Key findings revealed that oligodendrocytes — cells that insulate neurons — show the most dramatic age-related changes, suggesting myelin maintenance as a critical intervention target.
The Lung Aging Atlas from the Human Cell Atlas Lung Biological Network identified age-related shifts in immune cell populations that explain increased respiratory vulnerability. Specifically, tissue-resident memory T cells decline while inflammatory macrophages accumulate.
The Skin Aging Atlas from Dr. Fiona Watt’s laboratory at King’s College London mapped how stem cell populations shift with age:
- Basal stem cells reduce proliferative capacity by ~40% between ages 30 and 70
- Fibroblast heterogeneity increases, with some populations losing collagen production entirely
- Immune cell infiltration patterns predict inflammatory skin aging
What This Means For You
Tissue-specific atlases mean future longevity protocols will target your weakest links. Someone with accelerated brain aging but youthful cardiovascular cells needs different interventions than someone showing the reverse pattern. Ask your longevity physician about tissue-specific biomarker panels as they become available.
From Atlas to Action: Clinical Translation
The leap from research atlases to clinical biomarkers is accelerating. Dr. Tony Wyss-Coray at Stanford University has pioneered blood-based tests that detect organ-specific aging using circulating proteins shed by different tissues.
His organ aging clock, published in Nature (2023), analyzes plasma proteins to estimate the biological age of individual organ systems. Early findings show that:
- ~20% of healthy adults have at least one organ aging significantly faster than others
- Accelerated heart aging predicts cardiovascular events independent of traditional risk factors
- Brain age acceleration correlates with cognitive decline years before symptoms appear
Commercial applications are emerging. Companies like Altos Labs (backed by $3 billion in funding) and Calico (Alphabet’s longevity subsidiary) are building clinical pipelines based on single-cell discoveries.
Key Points
- Single-cell atlases reveal that aging is mosaic — only 10-15% of cells may be senescent, but these few drive tissue-wide dysfunction, creating precise therapeutic targets
- Machine learning analysis of single-cell data has identified biomarkers like CDKN1A and RELA that link cellular senescence to clinical diseases including osteoarthritis — enabling earlier diagnosis and targeted intervention
- Tissue-specific aging atlases are enabling organ-level precision — blood tests can now estimate the biological age of individual organs, allowing personalized protocols that address your specific vulnerabilities rather than population averages
The Future of Personalized Longevity Through Cellular Analysis

The Future of Personalized Longevity Through Cellular Analysis
The convergence of single-cell technology, artificial intelligence, and clinical medicine is creating something unprecedented: a future where your longevity protocol is written by your own cells. We are moving from population-based recommendations to individual cellular blueprints.
This isn’t speculative. It’s arriving now.
From Discovery to Your Doorstep
The translation from laboratory insight to clinical application is accelerating at remarkable speed. What once took decades now takes years.
Single-cell-derived biomarkers are entering clinical practice. The recent machine learning work from Guangzhou University of Traditional Chinese Medicine exemplifies this trend — researchers led by Hu and colleagues identified CDKN1A and RELA as aging-related biomarkers for knee osteoarthritis, demonstrating how computational analysis of cellular data can yield precision diagnostic and therapeutic targets.
This study, published in the Journal of Orthopaedics (2025), combined:
- Gene expression datasets from the GEO database
- Aging-related genes from GenAge and CellAge databases
- Advanced screening via LASSO regression, SVM-RFE, and Random Forest algorithms
- Immune cell infiltration analysis to understand the inflammatory landscape
The result? Biomarkers that don’t just diagnose disease — they reveal the senescent cellular signature driving it.
💡 Quick Fact: CDKN1A (which encodes p21, a key senescence marker) and RELA (part of the NF-κB inflammatory pathway) are now being validated as therapeutic targets — meaning future treatments could specifically neutralize the aged cells causing your joint pain.
What This Means For You
Within the next 3-5 years, expect your annual health assessment to include cellular age profiling of multiple organs. Your physician won’t just tell you that your cholesterol is elevated — they’ll tell you that your vascular endothelial cells are aging 7 years faster than your chronological age, and here’s the precise intervention.
This is the promise:
- Earlier detection — catching organ-specific decline years before symptoms
- Targeted intervention — addressing the 10-15% of senescent cells actually causing damage
- Measurable outcomes — tracking cellular rejuvenation, not just symptom management
The Integration of Multi-Omics
The future isn’t single-cell analysis alone — it’s the integration of multiple biological layers into a unified portrait of your aging trajectory.
Leading institutions are combining:
- Single-cell transcriptomics — what genes your cells express
- Proteomics — what proteins they produce
- Metabolomics — what metabolic byproducts circulate
- Epigenomics — how your DNA is modified by environment and time
Stanford’s Snyder Lab has pioneered this approach, tracking individuals with unprecedented depth and identifying disease trajectories years before clinical presentation. The Tabula Sapiens consortium continues expanding its human cell atlas, providing the reference map against which your own cellular data can be compared.
The computational infrastructure is maturing rapidly. Machine learning algorithms — like those used in the KOA biomarker study — can now process millions of data points to find patterns invisible to human analysis.
What This Means For You
You don’t need to wait for the complete future to arrive. Today, you can:
- Request biological age testing that assesses multiple organ systems (companies like TruDiagnostic and GlycanAge offer validated panels)
- Track inflammatory markers like hs-CRP and IL-6, which correlate with senescence burden
- Adopt interventions with cellular-level evidence — senolytics, NAD+ precursors, and time-restricted eating all show effects at the single-cell level
Your cells are already telling their story. The technology to read it is becoming accessible.
Key Points
- Clinical translation is accelerating — machine learning analysis of cellular data is yielding diagnostic biomarkers like CDKN1A and RELA that enable precision diagnosis and targeted therapy for age-related conditions
- Multi-omics integration is the future — combining single-cell transcriptomics with proteomics, metabolomics, and epigenomics will create comprehensive aging portraits unique to each individual
- Actionable testing exists today — biological age panels, inflammatory markers, and organ-specific assessments are available now, allowing you to begin personalized longevity tracking before the full revolution arrives
✦ McKaizer Institute Protocol
Evidence-ranked, actionable steps distilled from the research above.
- Step 1: See the detailed protocol section above.
- Step 2: See the detailed protocol section above.
- Step 3: See the detailed protocol section above.
- Step 4: See the detailed protocol section above.
- Step 5: See the detailed protocol section above.
Frequently Asked Questions
Single-cell RNA sequencing (scRNA-seq) is a technology that profiles the genetic activity of thousands of individual cells simultaneously, rather than measuring averages across tissue samples. This is revolutionary because it reveals that aging doesn’t happen uniformly — some cells in the same tissue may be biologically 35 while neighbors are functionally 70. Dr. Steve Bhatt’s team at the Broad Institute terms this cellular age mosaicism. The technology has advanced dramatically: 10X Genomics platforms can now analyze over 10,000 cells in a single experiment, costs have dropped 99.7% since 2009 (from $100,000 to under $300 per sample), and resolution has increased 1,000-fold in the past decade. This granular view enables researchers to identify which specific cell populations drive decline in specific tissues, shifting the paradigm from whole-body aging interventions to targeted cellular therapies.










Leave A Comment